Un interesante artículo de divulgación sobre la aparición del oxígeno en la atmósfera de nuestro planeta…
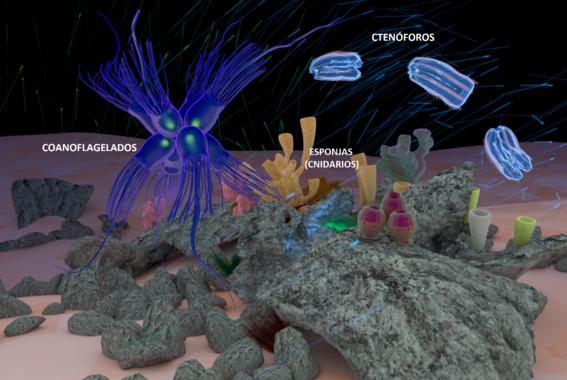
“Sabemos que en los primeros años de nuestro planeta el oxígeno estaba totalmente ausente.” “Pero si pudiéramos viajar a esta época -equipados, eso sí, con un buen balón de oxígeno-” … “Es probable que nos fijásemos más en que el mar es de color verde, y el cielo de un anaranjado turbio. No nos hemos equivocado de planeta, es que si cambiamos la química cambian los colores.” “La irrupción del oxígeno terminó con estas maravillas de la antigüedad.” … “a partir de los primeros años del paleoproterozoico se produce un aumento del oxígeno a escala global.” … “En las rocas, la proporción entre isótopos de azufre cambia, informándonos de que se ha formado una capa de ozono incipiente; los minerales, en general, aparecen más y más oxidados. En 300 millones de años el oxígeno se asienta, tanto en la atmósfera como en las aguas menos profundas de los océanos. A esta transición la llamamos la Gran Oxigenación, y es la prueba de que una nueva estirpe, las bacterias productoras de oxígeno, se estaba empezando a adueñar del mundo.” … “Gracias al oxígeno este mundo antiguo fue convirtiéndose en el que conocemos: el hierro de los océanos empezó a oxidarse y cayó rápidamente al fondo; por primera vez, los mares exhibieron un color azul que hoy nos es familiar: el color del agua.”